
Here we first demonstrated comparable sensitivity of the system to existing droplet-based methods by performing scRNA-seq on cell lines and synthetic RNAs. An analysis pipeline, Cell Ranger, processes the sequencing data and enables automated cell clustering. The resulting libraries then undergo Illumina short-read sequencing. Reverse transcription takes place inside each droplet, and barcoded complementary DNAs (cDNAs) are amplified in bulk. 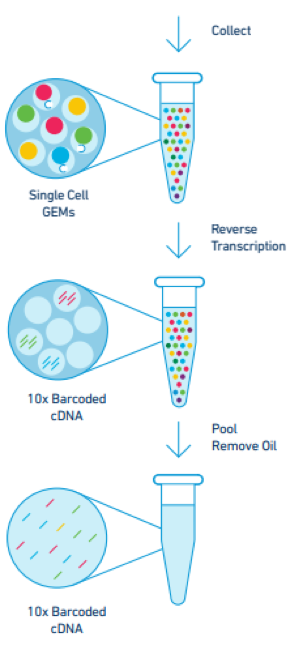
Approximately 50% of cells loaded into the system can be captured, and up to eight samples can be processed in parallel per run. To overcome these challenges, we developed a droplet-based system that enables 3′ messenger RNA (mRNA) digital counting of thousands of single cells. Droplet-based techniques have enabled processing of tens of thousands of cells in a single experiment 7, 8, but current approaches require generation of custom microfluidic devices and reagents. Plate-based methods often require time-consuming fluorescence-activated cell sorting (FACS) into many plates that must be processed separately 4, 9. Commercially available, microfluidic-based approaches have limited throughput 5, 6. However, previously described scRNA-seq methods face practical challenges when scaling to tens of thousands of cells or when it is necessary to capture as many cells as possible from a limited sample 4, 5, 6, 7, 8, 9. scRNA-seq studies have led to the discovery of novel cell types and provided insights into regulatory networks during development 3. Single-cell RNA-sequencing (scRNA-seq) can be used to dissect transcriptomic heterogeneity that is masked in population-averaged measurements 1, 2. Understanding of biological systems requires the knowledge of their individual components.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
